 Halford SE (1999) Restriction enzymes that act simultaneously at two DNA sites. Grazulis S, Deibert M, Rimseliene R, Skirgaila R, Sasnauskas G, Lagunavicius A, Repin V, Urbanke C, Huber R, Siksnys V (2002) Crystal structure of the Bse634I restriction endonuclease: comparison of two enzymes recognizing the same DNA sequence. Galburt EA, Stoddard BL (2002) Catalytic mechanisms of restriction and homing endonucleases. Nat Struct Biol 7:792–799Įmbleton ML, Siksnys V, Halford SE (2001) DNA cleavage reactions by Type II restriction enzymes that require two copies of their recognition sites. EMBO J 18:5805–5816ĭeibert M, Grazulis S, Sasnauskas G, Siksnys V, Huber R (2000) Structure of the tetra-meric restriction endonuclease NgoMIV in complex with cleaved DNA. J Mol Biol 255:176–186ĭeibert M, Grazulis S, Janulaitis A, Siksnys V, Huber R (1999) Crystal structure of MunI restriction endonuclease in complex with cognate DNA at 1.7 Åresolution. J Mol Biol 280:1–9īozic D, Grazulis S, Siksnys V, Huber R (1996) Crystal structure of Citrobacter freundii restriction endonuclease Cfr10I at 2.15 Å resolution. J Biol Chem 274:36379–36386īogan AA, Thorn KS (1998) Anatomy of hot spots in protein interfaces. Protein Sci 4:2455–2468īilcock DT, Daniels LE, Bath AJ, Halford SE (1999) Reactions of Type II restriction endonucleases with 8-base pair recognition sites. EMBO J 10:25–33īennett MJ, Schlunegger MP, Eisenberg D (1995) 3D domain swapping: a mechanism for oligomer assembly. J Biol Chem 277: 4024–4033īeese LS, Steitz TA (1991) Structural basis for the 3’–5’ exonuclease activity of Escherichia coli DNA polymerase I: a two metal ion mechanism. Curr Opin Struct Biol 5:11–19īath AJ, Milsom SE, Gormley NA, Halford SE (2002) Many Type IIs restriction endonucleases interact with two recognition sites before cleaving DNA. This process is experimental and the keywords may be updated as the learning algorithm improves.Īggarwal AK (1995) Structure and function of restriction endonucleases. These keywords were added by machine and not by the authors. Hence, the symmetry of the recognition sequence implies the oligomeric state of the restriction enzyme. Symmetrical nicking of opposite DNA strands by both monomers within a homodimer generates the double-strand break. Each subunit faces the same nucleotide sequence on the opposite DNA strand and contains one active site. 1), recognition of the palindromic DNA sequence is achieved by two identical protein subunits related by a twofold axis of symmetry. 
On the basis of this observation, Kelly and Smith proposed the first model for the interaction of these restriction enzymes with DNA ( Kelly and Smith 1970).According to their model (Fig. e., possess a twofold rotational axis of symmetry. Most of the sequences recognized by Type II restriction endonucleases are palindromic, i. 
Around 3500 species, from variety of bacteria with nearly 240 differing specificities, have now been characterized ( Roberts et al. 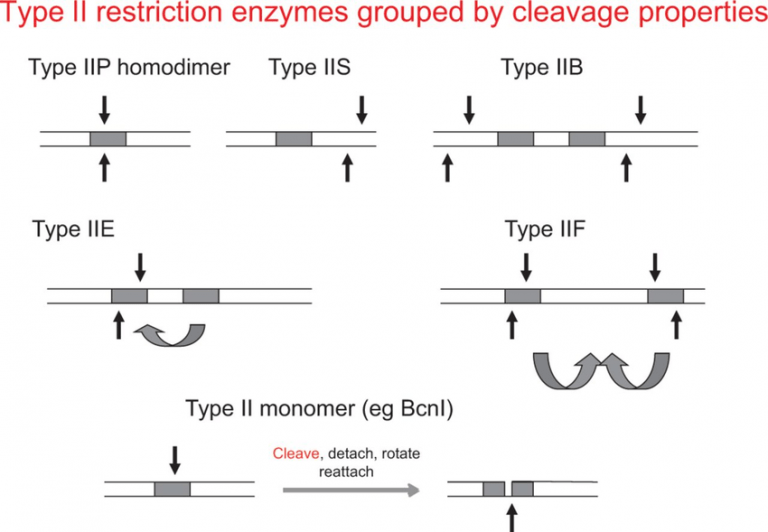
Type II restriction endonucleases recognize specific DNA sequences, typically 4–8 bp in length, and cleave phosphodiester bonds in the presence of Mg 2+, within or close to their recognition sites ( Pingoud and Jeltsch 2001). 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
